
MiR-129-5p: a novel therapeutic target for amyotrophic lateral sclerosis?
Amyotrophic lateral sclerosis (ALS) is a debilitating neurodegenerative disease that is defined by the progressive loss of motor function due to degeneration of upper and lower motor neurons in the brain and spinal cord tissue, respectively, with an average life-expectancy of 2–5 years post-diagnosis (1). Riluzole and edaravone are currently the only two FDA approved drugs for patients with ALS, but these drugs have minimal effect, extending life only for a matter of months (2).
In 1993, the first causative mutations of ALS were identified in the SOD1 gene (3). Today, mutations in SOD1 is the second most common cause of genetic ALS (~5–6% of all ALS cases), only falling behind the hexanucleotide repeat expansions observed in C9ORF72 (~12–14% of all ALS cases). However, the vast majority of ALS cases (~80%) have no known genetic background giving no clear indication on the causative factor that results in the disease development (1). In the last decade, miRNAs have been increasingly implicated in the pathogenesis of ALS due to the mass dysregulation of these molecules observed in ALS spinal cord and motor neurons (4-8).
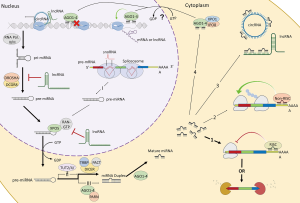
MiRNAs are small RNA molecules (~22–25 nucleotides) that are primarily responsible for post-transcriptional gene-silencing generally through interactions with the 3’ untranslated region (UTR) of messenger RNA (mRNA) (9). Their interactions are highly dependent on environmental cues. Therefore, cell development, cell type, cell stress, and aging all have an impact on mRNA post-transcriptional regulation via miRNAs. This is an especially true for neurons which have spatiotemporal needs within the cell where post-transcriptional regulation via miRNAs differs depending on soma, dendritic and axonal needs (9,10). Further, miRNAs are also regulated by other non-coding RNAs (i.e., long non-coding RNAs, circular RNAs, etc.) which work together to control the degree of expression of a gene (11), highlighting the sophistication of these non-coding RNA networks (Figure 1). Not surprisingly, because of miRNAs dynamic nature in these RNA networks, it has become increasingly of interest to determine the contribution of miRNAs in disease.
As mentioned, miRNAs have become increasingly implicated in ALS pathogenesis. This has mainly stemmed from observations that in sporadic ALS (sALS) spinal cord and motor neurons there are pools of miRNAs that are reduced (4-6). In contrast, mutant SOD1 (mtSOD1) ALS cases and rodent models have observed that miRNAs are generally upregulated, suggesting inhibition of these miRNAs may have therapeutic effects (7,12,13). In particular, high levels of miR-155 in mtSOD1 rodent models has been reported to promote neuroinflammation. Interestingly, inhibition of this miRNA via antisense oligonucleotides (ASOs) has been shown to decrease neuroinflammation and alleviate ALS-like phenotypes in rodent models (7). Others have used miRNAs to reduce SOD1 expression, and in turn reduce mtSOD1s toxic effects, by intrathecally delivering artificial miRNAs that were cloned into an Adeno-associated virus (AAV) and target the SOD1 transcript. Use of SOD1 targeting artificial miRNAs has been shown to have therapeutic effects in both rodent and macaque models (14,15). However, the effectiveness of theses therapeutics in humans is still unknown. This previous research provides evidence of the utility of miRNAs as potential therapeutic targets, and further, highlights the use of either ASOs or AAV-miRNAs as ways to develop therapeutics that will potentially treat neurodegeneration.
Recent work published by Loffreda et al., 2020 in Progress in Neurobiology highlights miR-129-5p as another upregulated miRNA in rodent models of mtSOD1 ALS which, if inhibited using an ASO, could have therapeutic effects for patients with ALS (16). The research group looked at mtSOD1 (SOD1 G93A) in vitro neuronal cell models in addition to in vivo mouse models and determined that mtSOD1 causes an increase in miR-129-5p levels. Further, they observed increased levels of miR-129-5p in peripheral blood mononuclear cell (PMBC) samples of sALS patients, suggesting that this phenomenon may be related to ALS more broadly rather than just those who contain mutations in SOD1. These researchers reported that increased levels of miR-129-5p in both their in vitro and in vivo models correlates with decreased levels of ELAVL4, otherwise known as HuD—a protein critical for neuronal cell identity, maturation, axogenesis, dendrite growth, plasticity and survival (17). Through robust experimental analyses, the authors were able to show that increased levels of miR-129-5p were directly responsible for the reduced levels of HuD, identifying the biological mechanism by which high levels of miR-129-5p may contribute to neurodegeneration. Having made this observation, the authors then examined the ability of an ASO targeting miR-129-5p to reduce its levels in mtSOD1 rodent models. Administration of the ASO via intracerebroventricular injection at an early symptomatic stage (post-natal 80 days) led to increased survival, improved motor capabilities, and increased neuromuscular junction connections. However, it only had a mild effect on preventing motor neuron degeneration which was not statistically significant. The authors concluded that while reducing levels of miR-129-5p may have some therapeutic effects, the most optimal approach may be to pair their ASO with another miRNA-based therapeutic currently in development—an ASO targeting miR-155 (7,18).
ASO-based therapeutics have become popular as an approach to treat neurodegenerative diseases due to the success in treating Spinal Muscular Atrophy (SMA)—a juvenile neurodegenerative disease caused by loss of lower motor neurons —using an ASO targeting SMN2 (19). Several modifications to the nucleotide backbone of an ASO can be done to increase its affinity to its target and make it resistant to nucleases. For example, morpholino modifications were used to develop the SMN2 ASO and the miR-129-5p ASO (16,19). Morpholino modifications alter the sugar-phosphate backbone by changing the sugar ring from a five-membered to a six-membered ring and the sugar rings are linked to phosphoramidates groups rather than phosphate groups. These modifications have been proven to be beneficial to develop ASOs that either knockdown expression of a specific gene, as seen in Loffreda et al., 2020, or alter mRNA splicing as seen for the ASO that targets SMN2 (16,19). Moreover, the ASO produced in Loffreda et al., 2020 is called an antagomir—an ASO that targets a miRNA—and they have been increasingly shown to be effective therapeutics. For example, the drug miravirsen, which has completed phase II clinical trials, is an antagomir targeting miR-122 and has been shown to prevent propagation of hepatitis C virus (HCV) RNA in the liver highlighting the potential use of antagomirs in clinical practice (20). Despite these developments, ASOs still have their limitations which include their potential off-target affects, their expense to develop, and their inability to cross the blood-brain barrier; making them difficult to administer for neurological diseases (21). However, the current success of ASO treatments thus far has sparked more innovative research to determine whether further modifications to these oligos will allow us to overcome these limitations, offering new opportunities for therapeutic development.
Loffreda et al., 2020 provides clear evidence that increased levels of miR-129-5p is caused by mtSOD1 and that lowering its levels in mtSOD1 mice has some therapeutic effects; however, it is unclear whether reduction of miR-129-5p would be a good ALS therapeutic more broadly (16). The authors identified increased levels of miR-129-5p in sALS patient PBMCs, but not in human spinal cord or motor neurons (16). As mentioned previously, the environmental context matters when discussing miRNA regulation (9). Further, in the several studies that have examined miRNA expression in the spinal cord and motor neurons of sALS cases, upregulation of miR-129-5p has not been observed (4-6). Thus, it is unclear whether this observation of increased levels of miR-129-5p in PBMCs is relevant to the dysfunction of motor neurons, or more broadly, the central nervous system of the sALS patient group within their study.
Further, the use of an ASO to suppress miR-129-5p expression should be approached with caution. In the central nervous system, miR-129-5p has been shown to suppress apoptosis and alleviate neuroinflammation (22). Low levels of miR-129-5p have been reported in the degenerating brain of patients with Alzheimer’s Disease, and lack of this miRNA creates an inflammatory response that can be neurotoxic (23). Thus, too much or too little of miR-129-5p could be damaging to the central nervous system. This is likely because miRNAs can target several transcripts to regulate a network of genes, which can make cells sensitive to fluctuations in miRNA expression. In this case, the authors showed that miR-129-5p regulates HuD levels, but miR-129-5p has also been shown to regulate critical neuronal genes involved in synaptic plasticity (FMR1) and neuroinflammation (HMGB1) (22,24). All this suggests that further work needs to be done to ensure reduction of this miRNA will not have long-term effects that may be more damaging than therapeutic.
Loffreda et al., 2020 proposed that while their ASO had some beneficial effects in their mtSOD1 models, matching it with an ASO that inhibits miR-155 may have the greatest overall therapeutic effect (16). This points to the idea that simply targeting one miRNA may not be enough to ameliorate ALS. In fact, as mentioned previously, several miRNAs have been shown to be upregulated in mtSOD1 models (7,12,13). Further, the article provides data indicating that mtSOD1 increased the levels of DICER, a critical protein in miRNA biogenesis, which may explain why miRNAs are upregulated in mtSOD1 models (16). Therefore, there seems to be a systemic issue with miRNA production in the presence mtSOD1. Thus, lowering levels of DICER might seem to be a better target to slow the production and reduce the toxicity of these upregulated miRNAs in mtSOD1 ALS cases. However, knockout of DICER in mouse motor neurons has been shown to lead to motor neuron degeneration, and thus, reducing levels of DICER should be approached with caution as well (25).
Overall, while this latest work by Loffreda et al., 2020 provides another interesting avenue for miRNA-based therapeutics for ALS, this work still has far to go before we can think about administering it to patients. Large-scale preclinical studies still need to be done to determine if inhibition of miR-129-5p prevents motor neurodegeneration, helps alleviate ALS-like phenotypes in several models of ALS or just SOD1-related models, and finally, determine whether downregulation of this miRNA has any potential negative long-term outcomes (i.e., chronic neuroinflammation). Further, we still do not know whether miR-129-5p is upregulated within the spinal cord or motor neurons of mtSOD1 ALS cases, which will be necessary to address to ensure development of this ASO as a therapeutic is not in vain. All this taken together, miR-129-5p could be a potential therapeutic for ALS, but in agreeance with the authors, it is unlikely that targeting this miRNA alone will be enough to completely halt the disease progression (16). However, the mere identification of this miRNA, its biological network and its potential relevance to disease will be incredibly important to our understanding of mechanisms associated with ALS progression, opening more avenues for potential therapeutics.
Acknowledgments
Funding: None.
Footnote
Provenance and Peer Review: This article was commissioned and reviewed by the Section Editor Mengli Chen, PhD (Jiangsu Province Hospital, Nanjing Medical University, Nanjing, China).
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/ncri-20-5). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Taylor JP, Brown RH Jr, Cleveland DW. Decoding ALS: from genes to mechanism. Nature 2016;539:197-206. [Crossref] [PubMed]
- Chiò A, Mazzini L, Mora G. Disease-modifying therapies in amyotrophic lateral sclerosis. Neuropharmacology 2020;167:107986 [Crossref] [PubMed]
- Rosen DR, Siddique T, Patterson D, et al. Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 1993;362:59-62. [Crossref] [PubMed]
- Emde A, Eitan C, Liou LL, et al. Dysregulated miRNA biogenesis downstream of cellular stress and ALS-causing mutations: a new mechanism for ALS. The EMBO journal 2015;34:2633-51. [Crossref] [PubMed]
- Reichenstein I, Eitan C, Diaz-Garcia S, et al. Human genetics and neuropathology suggest a link between miR-218 and amyotrophic lateral sclerosis pathophysiology. Sci Transl Med 2019;11:eaav5264 [Crossref] [PubMed]
- Campos-Melo D, Droppelmann CA, He Z, et al. Altered microRNA expression profile in Amyotrophic Lateral Sclerosis: a role in the regulation of NFL mRNA levels. Molecular brain 2013;6:26. [Crossref] [PubMed]
- Butovsky O, Jedrychowski MP, Cialic R, et al. Targeting miR-155 restores abnormal microglia and attenuates disease in SOD1 mice. Ann Neurol 2015;77:75-99. [Crossref] [PubMed]
- Williams AH, Valdez G, Moresi V, et al. MicroRNA-206 delays ALS progression and promotes regeneration of neuromuscular synapses in mice. Science 2009;326:1549-54. [Crossref] [PubMed]
- Hawley ZCE, Campos-Melo D, Droppelmann CA, et al. MotomiRs: miRNAs in Motor Neuron Function and Disease. Front Mol Neurosci 2017;10:127. [Crossref] [PubMed]
- Rotem N, Magen I, Ionescu A, et al. ALS Along the Axons - Expression of Coding and Noncoding RNA Differs in Axons of ALS models. Sci Rep 2017;7:44500. [Crossref] [PubMed]
- Salvatori B, Biscarini S, Morlando M. Non-coding RNAs in Nervous System Development and Disease. Front Cell Dev Biol 2020;8:273. [Crossref] [PubMed]
- Zhou F, Guan Y, Chen Y, et al. miRNA-9 expression is upregulated in the spinal cord of G93A-SOD1 transgenic mice. Int J Clin Exp Pathol 2013;6:1826-38. [PubMed]
- Marcuzzo S, Kapetis D, Mantegazza R, et al. Altered miRNA expression is associated with neuronal fate in G93A-SOD1 ependymal stem progenitor cells. Exp Neurol 2014;253:91-101. [Crossref] [PubMed]
- Stoica L, Todeasa SH, Cabrera GT, et al. Adeno-associated virus-delivered artificial microRNA extends survival and delays paralysis in an amyotrophic lateral sclerosis mouse model. Ann Neurol 2016;79:687-700. [Crossref] [PubMed]
- Borel F, Gernoux G, Sun H, et al. Safe and effective superoxide dismutase 1 silencing using artificial microRNA in macaques. Sci Transl Med 2018;10:eaau6414 [Crossref] [PubMed]
- Loffreda A, Nizzardo M, Arosio A, et al. miR-129-5p: A key factor and therapeutic target in amyotrophic lateral sclerosis. Prog Neurobiol 2020;190:101803 [Crossref] [PubMed]
- Bronicki LM, Jasmin BJ. Emerging complexity of the HuD/ELAVl4 gene; implications for neuronal development, function, and dysfunction. RNA 2013;19:1019-37. [Crossref] [PubMed]
- Koval ED, Shaner C, Zhang P, et al. Method for widespread microRNA-155 inhibition prolongs survival in ALS-model mice. Hum Mol Genet 2013;22:4127-35. [Crossref] [PubMed]
- Finkel RS, Mercuri E, Darras BT, et al. Nusinersen versus Sham Control in Infantile-Onset Spinal Muscular Atrophy. N Engl J Med 2017;377:1723-32. [Crossref] [PubMed]
- Janssen HL, Reesink HW, Lawitz EJ, et al. Treatment of HCV infection by targeting microRNA. N Engl J Med 2013;368:1685-94. [Crossref] [PubMed]
- Schoch KM, Miller TM. Antisense Oligonucleotides: Translation from Mouse Models to Human Neurodegenerative Diseases. Neuron 2017;94:1056-70. [Crossref] [PubMed]
- Wan G, An Y, Tao J, et al. MicroRNA-129-5p alleviates spinal cord injury in mice via suppressing the apoptosis and inflammatory response through HMGB1/TLR4/NF-kappaB pathway. Biosci Rep 2020;40:BSR20193315 [Crossref] [PubMed]
- Zeng Z, Liu Y, Zheng W, et al. MicroRNA-129-5p alleviates nerve injury and inflammatory response of Alzheimer's disease via downregulating SOX6. Cell Cycle 2019;18:3095-110. [Crossref] [PubMed]
- Zongaro S, Hukema R, D'Antoni S, et al. The 3' UTR of FMR1 mRNA is a target of miR-101, miR-129-5p and miR-221: implications for the molecular pathology of FXTAS at the synapse. Hum Mol Genet 2013;22:1971-82. [Crossref] [PubMed]
- Haramati S, Chapnik E, Sztainberg Y, et al. miRNA malfunction causes spinal motor neuron disease. Proc Natl Acad Sci U S A 2010;107:13111-6. [Crossref] [PubMed]
Cite this article as: Hawley ZCE, Campos-Melo D, Strong MJ. MiR-129-5p: a novel therapeutic target for amyotrophic lateral sclerosis? Non-coding RNA Investig 2020;4:7.


