Neonatal rat fibroblast isolation, culture, differentiation and determination of microRNA effects in transformation
Introduction
Cardiac fibrosis is characterized by extracellular matrix (ECM) accumulation in the cardiac interstitium and leads to both systolic and diastolic dysfunction in many pathological conditions, not only in myocardial infarction, but also in the volume and pressure overload myocardium, in alcoholic cardiomyopathy and in the aging heart (1-3). Treatments for fibrosis are usually done by controlling inflammation and protecting from any deterioration of the cardiac function (4). Recent studies have demonstrated that although angiotensin II (AngII) receptor antagonists and ACE inhibition can reverse cardiac remodeling (5,6), there are no effective anti-fibrotic treatments available. Currently, mounting data have suggested that microRNAs (miRNA) could lead to a profound regulation of target genes and related signaling pathways, engaging in many pathological conditions including adverse cardiac remodeling, heart failure and fibrosis (7-9), which raise a hope that manipulating miRNAs may provide a novel potential treatment to combat cardiac fibrosis.
Quiescent cardiac fibroblasts are the most abundant cells in the unstressed mammalian heart which have no stress fibers or contractile microfilaments, processing few actin-associated cell-cell or cell-matrix contacts and do not secrete amounts of matrix proteins (10,11). In the aftermath of damaged myocardium, fibroblasts are activated, proliferate and differentiate into myofibroblasts, which are the dominant effector cells in the fibrotic ventricle and play key roles through structural, paracrine, and electrical links with cardiomyocytes (12). Myofibroblasts exhibit stellate forms and feature stress fibers with highly developed focal adhesions. Fully differentiated myofibroblasts express α-smooth muscle actin (α-SMA) and other smooth muscle cells (SMC) markers, they are able to secrete ECM components and synthesize collagen, contributing to wound healing, matrix remodeling and eventual cardiac fibrosis through the elevated production of collagens (13,14).
Myofibroblasts transdifferentiation is a hallmark of the cardiac fibrotic response (15). A growing body of evidence suggests that in vitro, primary neonatal or adult ventricular passage 0 (P0) fibroblasts experience rapid and consistent transdifferentiation to myofibroblasts upon standard culture conditions (using Dulbecco’s Modified Eagle Medium-DMEM based culture media supplemented with 10% fetal bovine serum) (16). In addition, myofibroblasts are highly responsive to several cytokines, chemokines and growth factors in serum free conditions, which initiate myofibroblast differentiation. Transforming growth factor-β (TGF-β) has been identified as a primary and component mediator of myofibroblasts transdifferentiation. The introduction of TGF-β thus appears to be a universal paradigm for the transformation of isolated fibroblasts in vitro. AngII is a member of the renin-angiotensin system which appears to dominantly affect the cardiac fibrotic response and myofibroblast activation (17-20). Therefore, exposure of fibroblasts to AngII is another model of myofibroblast transdifferentiation.
Currently no protocol has been systematically provided for neonatal fibroblast isolation, culture, transdifferentiation and miRNA transfection. Here we provided a detail protocol for fibroblast isolation, culture, transformation and miRNA transfection. As cardiac fibrosis has profound consequences on myocardial function, this protocol may have extremely importance for developing novel therapeutics for cardiac fibrosis.
Materials
Tissue samples: Neonatal rats (0–3-day old) can be purchased from Sprague-Dawley (SD) colonies (see Note I). All the materials and solutions used in cell manipulation should be sterilized either by irradiation or 0.22 µm filtering, these materials should also be disposable cell culture grade (see Note II). Surgical scissors, tweezers and other instruments were sterilized by autoclaving in advance before starting. All regents used are stored at room temperature (20–23 °C) unless indicated.
Fibroblast isolation and culture
- 10× ADS solution (100 mL): 4.76 g HEPES, 6.8 g Nacl (Sigma-Aldrich) + 0.138g Na2HPO4 (Sigma-Aldrich) +0.6 g glucose (Sigma-Aldrich) +0.4 g Kcl (Sigma-Aldrich) +0.051 g MgSO4 (containing 7H2O, Sigma-Aldrich), dissolved to 100 mL with ultrapure water, adjust pH value to 7.35–7.45 and finally filtrate with 0.22 µm filter. Store at 4 °C (see Note III).
- 1× ADS solution (1 L): 100 mL 10× ADS + 900 mL sterile water. Store at 4 °C.
- Trypsin-collagenase 2 buffer (100 mL for heart dissociations): 60 mg trypsin (Sigma-Aldrich) +40 mg collagenase (Worthington), dissolved to 100 mL with 1× ADS solution. Store at 4 °C. Incubate at 37 °C for 1–2 h before use.
- Horse serum (HS, Gibco) and fetal bovine serum (FBS, Gibco): store at −20 °C. Incubate at 37 °C for 1 h before use.
- Mixture of penicillin (10,000 U/mL, Hyclone) and streptomycin (10,000 µg/mL, Hyclone): store at −20 °C. Incubate at 37 °C for 30 min before use.
- Culture medium for neonatal rat fibroblasts (completed DMEM 500 mL): 450 mL DMEM (Hyclone) + 50 mL FBS + 6 mL Penicillin-Streptomycin solution (P/S). Incubate at 37 °C for 30 min before use.
Myofibroblast transdifferentiation
- Phosphate buffer saline (PBS, Gibco): store at 4 °C.
- 0.25% Trypsin + EDTA (Gibco): store at −20 °C.
- Human/rhesus/canine TGF-beta 1/TGFB1 protein (10 ng/mL, Sino Biological Inc., 10804-HNAC): store at −20 °C.
- Angiotensin II human (100 nmol/L, Sigma-Aldrich, A9525): store at −20 °C (see note IV).
- Serum-free culture medium for myofibroblast (serum-free DMEM 500 mL): 494 mL DMEM +6 mL P/S. Incubate at 37 °C for 30 min before use.
miRNA agomir/antagomir transfection
- miRNA agomir/antagomir/negative control (NC) stock solution (20 nmol, Ribobio): 5 nmol agomir/antagomir + 250 mL sterile water. Store at −20 °C.
- Reduced serum medium (Opti-MEM, Gibco): store at 4 °C.
α-SMA immunostaining
- Anti-Actin, a-Smooth Muscle-Cy3TM antibody, Mouse monoclonal (Sigma-Aldrich, C6198). Store at 4 °C.
- Fixing solution [4% paraformaldehyde (PFA), 100 mL, Bioshorp]: 4 mL PFA solution + 96 mL 1× PBS. Store at room temperature.
- Permeabilized solution (0.5% PBST, 100 mL, Bioshorp): 500 µL Triton-x-100 solution + 100 mL 1× PBS. Store at room temperature.
- Blocking solution (10% goat serum, 100 mL, Sigma-Aldrich): 10 mL goat serum + 90 mL 0.5% PBST. Store at 4 °C.
- 4', 6-Diamidino-2-Phenylindole (DAPI, Invitrogen). Store at −20 °C.
Methods
All procedures must comply with and have specific approval of your institution's Institutional Review Board. All the protocols were carried out at room temperature unless indicated.
Fibroblast isolation and culture
- Sacrifice neonates by IACUC-approved cervical dislocation.
- Small dissection scissors are used to cut through ribs and then the thoracic cavity is opened.
- Squeeze gently to expose the heart and then cardiac ventricle tissues are removed and wash several times in ice-cold 1× ADS (see Note V).
- Rat ventricle tissues are cut into pieces of approximately 1 mm3 and transfer to 1× ADS dish. When all hearts are removed to 1× ADS dishes, transfer the dishes to tissue culture hood.
- A transfer pipette is used to remove most of the 1× ADS from culture hood (see Note VI).
- Sterile forceps are used to transfer a group of ~15 ventricle tissues to sterile 1× ADS-wetted glass dish (see Note VII).
- Add 30 mL preheated trypsin-collagenase 2 buffer to each glass dish.
- Stir in preheated cell vibrator at 100 rpm, 37 °C for 30 min. Do not let the temperature exceed 37 °C.
- Carefully transfer the supernatant to a 50-mL centrifuge tube (collecting tube) (see Note VIII) and add 6mL HS to the collecting tube to terminate enzyme activity.
- Centrifuge the collecting tube in a tabletop centrifuge at 1000 rpm for 5 min.
- Gently remove the supernatants and the remaining cell pellet is resuspended in 2 mL warmed neonatal fibroblast completed DMEM medium.
- Add fresh 30 mL preheated trypsin-collagenase 2 buffer to the remaining tissue pieces and stir in cell vibrator at 100 rpm, 37 °C for 20 min (see Note IX).
- Repeat step VIII–XIII until ventricular tissue fragments are completely dissolved (see Note X).
- Gently pool cell suspension and filter by 0.22 µm filtering into a new sterile 50 mL centrifuge tube (see Note XI).
- Centrifuge the 50-mL centrifuge tube in a tabletop centrifuge at 1000 rpm for 5 min.
- Gently remove the supernatants and suspend the remaining cell pellet in 10 mL warmed neonatal fibroblast completed DMEM medium.
- After fully mix the cell suspension, absorb 10 µL suspension and use a hemocytometer to count cells which are not red blood cells (RBC) (see Note XII).
- Calculate the number of cell clusters and add corresponding completed DMEM at ~4–5×106 cells/10 cm culture dish (see Note XIII).
- Dispense cell clusters in 10 cm culture dish. Rinse suspension tube with very small volume completed DMEM and add to already-plated dish.
- Incubate for 3 h in a 37 °C, 5% CO2 incubator to allow fibroblasts to adhere (see Note XIV).
- Use a transfer pipette to remove and discard media from all culture dishes, rinsing the remaining culture dish with overlying completed DMEM (see Note XV).
- Repeat step XXI for 3–4 times.
- Add 10 mL completed DMEM to each 10cm culture dish and fibroblasts (P0) are incubated in a 37 °C, CO2 incubator (see Note XVI, Figure 1).
Myofibroblast transdifferentiation by serum-based protocol
- Use a transfer pipette to remove and discard media from all culture dishes.
- Add 3–4 mL PBS to each culture dish.
- Use a transfer pipette to remove and discard PBS from all culture dishes.
- Add 1 mL 0.25% trypsin to each dish.
- Incubate in a 37 °C, 5% CO2 incubator for ~5–8 min.
- Remove clusters and tap sides to release cells.
- Check culture dishes by microscope to ensure cells are rounded and floating. If not, incubate several minutes longer (see Note XVII).
- Add 3–4 mL completed DMEM to each dish to determinate digestion.
- Use overlying media to rinse wells and pool in 50 mL centrifuge tube.
- Centrifuge in a tabletop centrifuge at 1,000 rpm for 5 min.
- Decant supernatants into waste and use 10 mL completed DMEM to resuspend the pellet.
- Calculate the number of cell clusters and add corresponding completed DMEM at a density of ~2–4×105 cells/mL (see Note XVIII).
- Add 10 mL completed DMEM to each 10 cm culture dish and fibroblasts (P1) are incubated in a 37 °C, CO2 incubator (see Note XIX).
- Repeat steps I–XIII to obtain P2 and P3 fibroblasts.
- The transformation of fibroblast to myofibroblast is detected by immunofluorescence of α-SMA.
Myofibroblast transdifferentiation by serum-free protocol (induced by TGF-β and AngII)
- Incubate fibroblast cultures (P2) 24 h and assess by light microscopy (see Note XX).
- Remove overlying media.
- Add corresponding volume of serum-free DMEM to each well and incubate for 6–8 h.
- Prepare TGF-β (10 ng/mL) or AngII (100 nmol/L) with serum-free DMEM.
- Remove overlying serum-free DMEM.
- Add TGF-β (10 ng/mL) or AngII (100 umol/L) to each well (see Note XXI).
- Incubate cells in a humidified incubator containing 5% CO2 at 37 °C. Fibroblasts will be treated for 24 h (TGF-β) or 48 h (AngII) (see Note XXII).
- At this point the cells can be transfected, or stained and then stored at room temperature in the dark (Figure 2).
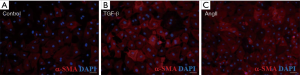 Figure 2 Transformation of fibroblast to myofibroblast stimulated by TGF-β and AngII. (A) α-SMA (red)/DAPI (blue) immunofluorescence staining of passage 2 (P2) fibroblast under serum-free DMEM media; (B) α-SMA (red)/DAPI (blue) immunofluorescence staining of P2 fibroblast stimulated by TGF-β; (C) α-SMA (red)/DAPI (blue) immunofluorescence staining of P2 fibroblast stimulated by AngII. Scale bar represented 50 µm. SMA, smooth muscle actin; DAPI, 4', 6-Diamidino-2-Phenylindole; TGF, transforming growth factor; AngII, angiotensin II.
Figure 2 Transformation of fibroblast to myofibroblast stimulated by TGF-β and AngII. (A) α-SMA (red)/DAPI (blue) immunofluorescence staining of passage 2 (P2) fibroblast under serum-free DMEM media; (B) α-SMA (red)/DAPI (blue) immunofluorescence staining of P2 fibroblast stimulated by TGF-β; (C) α-SMA (red)/DAPI (blue) immunofluorescence staining of P2 fibroblast stimulated by AngII. Scale bar represented 50 µm. SMA, smooth muscle actin; DAPI, 4', 6-Diamidino-2-Phenylindole; TGF, transforming growth factor; AngII, angiotensin II.
miRNA agomir/antagomir transfection
- Incubate fibroblast cultures (P2) 24 h and assess by light microscopy (see Note XXIII).
- Remove overlying media.
- Add corresponding volume of serum-free DMEM to each well and incubate for 6–8 h.
- Prepare agomir/antagomir and NC reaction mixture (Table 1).
Table 1
Working concentration of miRNA agomir/antagomir on different vesselsVessel Concentration of agomir (nM) Volume of agomir (ìL) Concentration of antagomir (nM) Volume of antagomir (ìL) 96 well 50 0.5 100 1 24 well 50 2.5 100 5 12 well 50 5 100 10 6 well 50 10 100 20 - Remove half of the overlying serum-free DMEM from each cell plate and add transfection mixture (see Note XXIV).
- Incubate cells in a humidified incubator containing 5% CO2 at 37 °C for 48 h.
- Assessing miRNA transfection efficiency: after 48 h post-transfection, fibroblast cultures are suitable for quantitation of: generation of miRNA RNA (qRT-PCR), miRNA-induced fibrotic effects (immunofluorescence, qRT-PCR or immunoblot).
α-SMA immunostaining
- Assess fibroblasts by light microscopy and then carefully aspirate culture media and wash once with PBS (24 well plate).
- Add 200 µL 4% PFA to each well and incubate for 20 min at room temperature.
- Carefully remove paraformaldehyde and wash three times with ice-cold PBS (see Note IIV).
- Add 200 µL 0.5% PBST to each well and incubate for 20 min at room temperature.
- Carefully remove PBST and wash three times with ice-cold PBS.
- Add 100 µL 10% goat serum to each well and incubate for 1 h at room temperature.
- Carefully remove 10% goat serum.
- Add 50 µL primary antibody: monoclonal alpha-smooth muscle actin (diluted in 1:100 in 10% goat serum) to each well and incubate overnight at 4 °C protected from light (see Note XXVI).
- Carefully remove primary antibody and wash three times with ice-cold PBS.
- Add 100 µL DAPI (Invitrogen, 1:10,000, diluted in PBS) to each well and incubate for 20 min at room temperature protected from light.
- Carefully remove DAPI mixture and wash three times with ice-cold PBS.
- Add 50 uL PBS to each well and protect from light.
- Examine wells with an upright metallurgical confocal (Zeiss) using appropriate filters (see Note XXVII).
Notes
- All tissue samples from 0–3-day old SD rats must be approved in advance by the appropriate Institutional Animal Care and Use Committee (IACUC).
- Cardiac fibroblasts are extraordinarily sensitive to contaminating residues induced by autoclaves. Therefore, during the process of isolation and culture of fibroblast, use disposable plastics rather than glasses if possible. If must be used, standard laboratory glassware should avoid exposing to endotoxins, detergents or other contaminants.
- The ADS, Trypsin-collagenase 2 buffer should be freshly prepared before use.
- When preparing AngII solution, pay attention to be away from light.
- After ventricle tissues are taken out from the body, irrelevant tissues such as blood vessels, atria and lung are needed to be removed. Then the ventricle tissues are washed in ice-cold 1× ADS several times to remove blood.
- This step could be repeated several times to fascinate access to hearts.
- In general, 15 ventricle tissues could be fully dissociated with 30 mL trypsin-collagenase 2 buffer.
- Before transferred the supernatants to centrifuge tubes, standing the glass dish for 5–10 min to obtain fibroblasts as far as possible.
- During the process of trypsinizations, appropriate reduce the volume of trypsin-collagenase 2 buffer and dissolved time according the amount of the remaining tissue fragments.
- Note the cloudiness of the supernatants each time, after peaking it would be decrease. In later trypsinizations, the supernatant might become “gooey”. Generally, eight trypsinizations were sufficient for full digestion of 15 rat hearts. If there was excessive “goo” substances, stop digestion earlier.
- It is highly advisable to filter cell suspension to remove collagen.
- When calculating cells, noting that RBCs are smaller, extremely round in shape, and have a “donut-like” appearance. Please do not count RBCs. Count at least 100 cells per loading.
- There should be 4–10×105 cells (non-RBC) per heart. For 15 neonatal rat hearts, there should be 60–150×105 cells (non-RBC). Generally, the total volume of cell suspension is 20 mL/15 rat hearts.
- By eye, at the end of incubation, the dishes would like they have a thick monolayer that starts to rip as you swirl the cluster. Cardiomyocytes are just aggregated and not adherent at the stage.
- When removing the cell suspension, leaving ~1 mL in the culture dishes and rocking the dishes each removal to ensure that cells do not dry out.
- Before passaging cells, fibroblasts cultures (P0) are left undisturbed for 36 hours.
- It may take a little longer to fully digest fibroblasts, slightly tap sides to release cells.
- The density has an effect on fibroblast differentiation, and the density of fibroblast cultured between 2–4×105 cells/mL is optimal for fibroblast differentiation. In addition, the density of cells varies according to different experimental needs (Table 2).
Table 2
Different cell density for different applicationVessel Volume of cell suspension (ìL/well) Cell number Example of purpose 24 well 500 1×105 Immunofluorescence staining 12 well 1,000 3×105 RNA extraction 6 well 2,000 8×105 Protein extraction (small) 6 cm culture dish 4,000 16×105 Protein extraction (large) - It is difficult to visualize cardiac fibroblasts by light microscopy because the cell borders are difficult to discern. Lowering the brightness of light may help to increase the contrast and visibility.
- As reported in our study, P2 fibroblasts are chosen to be stimulated by TGF-β or AngII (21).
- When preparing and dealing with AngII, pay attention to be away from light. It is better to be wrapped with foil before putting culture plates into incubator.
- After 48 h, fibroblasts will be obvious on a low power inverted microscope, due to their elongate morphology (50–200 µm long spindle-shaped cells).
- It is better to keep the cell density to 70–80% before miRNA transfection.
- To minimize the potential chemical toxicity of miRNA agomir/antagomir, it is better to remove half of the overlying serum-free DMEM from each cell plate before transfection.
- Paraformaldehyde should be carefully disposed as hazardous waste.
- As primary antibody monoclonal alpha-smooth muscle actin is RFP conjunct, there is no need for second antibody to incubate.
- 15–20 fields (200× magnification) were taken from every sample and the fluorescence gray value of α-SMA was calculated.
Acknowledgments
Funding: This work was supported by the grants from National Natural Science Foundation of China (81700343 to Lichan Tao and 81700351 to Hui Wang) and Natural Science Foundation of Jiangsu Province of China (BK20170296 to Lichan Tao).
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/ncri.2017.11.03). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All tissue samples have been approved in advance by the appropriate Institutional Animal Care and Use Committee (IACUC).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Leask A. Getting to the heart of the matter: new insights into cardiac fibrosis. Circ Res 2015;116:1269-76. [Crossref] [PubMed]
- Thum T. Noncoding RNAs and myocardial fibrosis. Nat Rev Cardiol 2014;11:655-63. [Crossref] [PubMed]
- Tao H, Shi KH, Yang JJ, et al. Epigenetic regulation of cardiac fibrosis. Cell Signal 2013;25:1932-8. [Crossref] [PubMed]
- Roubille F, Busseuil D, Merlet N, et al. Investigational drugs targeting cardiac fibrosis. Expert Rev Cardiovasc Ther 2014;12:111-25. [Crossref] [PubMed]
- Brilla CG, Rupp H, Maisch B. Effects of ACE inhibition versus non-ACE inhibitor antihypertensive treatment on myocardial fibrosis in patients with arterial hypertension. Retrospective analysis of 120 patients with left ventricular endomyocardial biopsies. Herz 2003;28:744-53. [Crossref] [PubMed]
- Elnakish MT, Kuppusamy P, Khan M. Stem cell transplantation as a therapy for cardiac fibrosis. J Pathol 2013;229:347-54. [Crossref] [PubMed]
- Creemers EE, van Rooij E. Function and Therapeutic Potential of Noncoding RNAs in Cardiac Fibrosis. Circ Res 2016;118:108-18. [Crossref] [PubMed]
- Bauersachs J. Regulation of myocardial fibrosis by MicroRNAs. J Cardiovasc Pharmacol 2010;56:454-9. [Crossref] [PubMed]
- Dai Y, Khaidakov M, Wang X, et al. MicroRNAs involved in the regulation of postischemic cardiac fibrosis. Hypertension 2013;61:751-6. [Crossref] [PubMed]
- Lajiness JD, Conway SJ. Origin, development, and differentiation of cardiac fibroblasts. J Mol Cell Cardiol 2014;70:2-8. [Crossref] [PubMed]
- Lighthouse JK, Small EM. Transcriptional control of cardiac fibroblast plasticity. J Mol Cell Cardiol 2016;91:52-60. [Crossref] [PubMed]
- Deb A, Ubil E. Cardiac fibroblast in development and wound healing. J Mol Cell Cardiol 2014;70:47-55. [Crossref] [PubMed]
- Kong P, Christia P, Frangogiannis NG. The pathogenesis of cardiac fibrosis. Cell Mol Life Sci 2014;71:549-74. [Crossref] [PubMed]
- Driesen RB, Nagaraju CK, Abi-Char J, et al. Reversible and irreversible differentiation of cardiac fibroblasts. Cardiovasc Res 2014;101:411-22. [Crossref] [PubMed]
- van Putten S, Shafieyan Y, Hinz B. Mechanical control of cardiac myofibroblasts. J Mol Cell Cardiol 2016;93:133-42. [Crossref] [PubMed]
- Santiago JJ, Dangerfield AL, Rattan SG, et al. Cardiac fibroblast to myofibroblast differentiation in vivo and in vitro: expression of focal adhesion components in neonatal and adult rat ventricular myofibroblasts. Dev Dyn 2010;239:1573-84. [Crossref] [PubMed]
- Davis J, Molkentin JD. Myofibroblasts: trust your heart and let fate decide. J Mol Cell Cardiol 2014;70:9-18. [Crossref] [PubMed]
- Chen C, Li R, Ross RS, et al. Integrins and integrin-related proteins in cardiac fibrosis. J Mol Cell Cardiol 2016;93:162-74. [Crossref] [PubMed]
- Wang YS, Li SH, Guo J, et al. Role of miR-145 in cardiac myofibroblast differentiation. J Mol Cell Cardiol 2014;66:94-105. [Crossref] [PubMed]
- Bai J, Zhang N, Hua Y, et al. Metformin inhibits angiotensin II-induced differentiation of cardiac fibroblasts into myofibroblasts. PLoS One 2013;8:e72120 [Crossref] [PubMed]
- Tao L, Bei Y, Chen P, et al. Crucial Role of miR-433 in Regulating Cardiac Fibrosis. Theranostics 2016;6:2068-83. [Crossref] [PubMed]
Cite this article as: Tao L, Wang H, Yang X. Neonatal rat fibroblast isolation, culture, differentiation and determination of microRNA effects in transformation. Non-coding RNA Investig 2017;1:18.